9. Clustering#
9.1. Practical Session 2: Deterministic Annealing#
9.1.1. Introduction#
In this session, we will explore the concept of deterministic annealing, a technique used for clustering data points into groups. Deterministic annealing is an optimization algorithm that extends the idea of simulated annealing to clustering problems. It allows us to find the optimal clustering of data points by gradually reducing the temperature parameter.
9.1.2. Deterministic Annealing#
Deterministic Annealing (DA) is a search technique easily applicable to the central clustering problem (see more details in Keneth Rose’s PhD Thesis).
The basic idea of DA is that for each value of \(\beta=\frac{1}{T}\) we interate two phases for minimizing the free energy \(F_{\beta}({\cal C})\):
Expectation. For fixed centers \(\mathbf{c}_i\), we estimate the probability that any point \(\mathbf{x}_a\) “belongs” to each cluster (“membership”). We denote such probabilities as \(\langle \mathbf{M}_{ai} \rangle\) and they are given by:
Update. For fixed probabilities \(\langle \mathbf{M}_{ai} \rangle\), we update the centers. If \(D(\mathbf{x}_a,\mathbf{c}_i)=||\mathbf{x}_q-\mathbf{c}_i||^2\) (squared Euclidean distance), then we have
i.e. the new (“fuzzy”) centers are the expected centers according to the fixed probabilities or (“memberships”).
Algorithm 9.1 (Deterministic Annealing [Clustering])
Inputs Given a set of points \({\cal X}=\{\mathbf{x}_1,\ldots,\mathbf{x}_m\}\), the free energy \(F_{\beta}({\cal C})\), an annealing schedule \(\beta =1/T(t)\) and an inital state \({\cal C}^0=(\mathbf{c}^0_1,\ldots,\mathbf{c}^0_k)\)
Outputs \({\cal C}^{\ast}=\arg\min_{{\cal C}\in\Omega} F_{\beta\rightarrow\infty}({\cal C})\) and \(\langle \mathbf{M}^{\ast}_{ai}\rangle\)
convegence = False
\(\langle \mathbf{M}^{old}_{ai}\rangle\leftarrow 1\)
\(t\leftarrow 0\)
while \(\neg\) convergence:
for \(\mathbf{x}_a\in {\cal X}\), \(\mathbf{c}_i\in {\cal C}\):
\( \langle \mathbf{M}_{ai} \rangle = \frac{\exp(-\beta D(\mathbf{x}_a,\mathbf{c}_i))}{\sum_{i'=1}^k \exp(-\beta D(\mathbf{x}_a,\mathbf{c}_{i'}))} \)
for \(\mathbf{c}_i\in {\cal C}\):
\( \mathbf{c}_i = \frac{\sum_{a=1}^m \mathbf{x}_a \langle \mathbf{M}_{ai}\rangle}{\sum_{a=1}^m\langle \mathbf{M}_{ai}\rangle} \)
\(t\leftarrow t+1\)
\(T(t)\leftarrow \frac{1}{\log(1 + t)}\)
convergence = \((T(t)<T_{min})\) or \((t>t_{max})\) or \((\sum_{ai}|\langle \mathbf{M}_{ai}\rangle - \langle \mathbf{M}^{old}_{ai}\rangle|\le\epsilon )\)
\(\langle \mathbf{M}^{old}_{ai}\rangle\leftarrow \langle \mathbf{M}_{ai}\rangle\)
return \({\cal C}^{\ast}\), \(\langle M^{\ast}\rangle\)
Some considerations about the above algorithm:
It is deterministic, i.e. we do not draw random numbers at any step of the algoorithm.
The initial (inverse) temperature \(\beta=1/T_{max}\), where \(T_{max}=T(0)\), is used as in SA, to make all the assignments almost equally probable: note that \(e^{-\beta D(\mathbf{x}_a,\mathbf{c}_i)}\rightarrow 1\), if \(\beta\rightarrow 0\). Under these conditions, we have \(\langle M_{ai}\rangle\approx 1/k\).
Initialization. \(T_{max}\) is set to the maximum variance \(\sigma^2_{max}\) of the data. If the data is multidimensional, as it happens usually, \(\sigma^2_{max}\) has an spectral interpretation (PCA).
However, as the algorithm evolves, \(\beta\) increases and the exponential decay becomes more selective: given \(D(\mathbf{x}_a,\mathbf{c}_i)\le D(\mathbf{x}_a,\mathbf{c}_j)+\alpha\), the second distance decays exponentially faster than the first as \(\beta\) increases.
Alternating Expectation and Update. Given the probabilities \(\langle M_{ai}\rangle\) we update the centers \(\mathbf{c}_i\) and then we re-compute the probabilities until a “fixed point” (stable assignment/centers) is reached.
Convervence. The algorithm converges to the nearest local optimum of the free energy to the initialization point \({\cal C}^0\). We add the condition \(\sum_{ai}|\langle \mathbf{M}_{ai}\rangle - \langle \mathbf{M}^{old}_{ai}\rangle|\le\epsilon\) with \(\epsilon>0\), which means that the algorithm stops if the \(\langle \mathbf{M}_{ai}\rangle\) are stable enough. For instance, if we start by setting \(\langle M_{ai}\rangle\approx 1/k\), the algorithm only performs a single iteration. Why? Because the centers in step 4.2. are not modified at all.
Complexity. Each iteration takes \(O(mk)\) and the number of iterations can be accelerated by a faster annealing schedule.
Outputs. The algorithm returns the best centers \({\cal C}^{\ast}\) and the optimal assignments \(\langle \mathbf{M}_{ai}^{\ast}\rangle\) where a point \(\mathbf{x}_a\) is assigned to the cluster centered at \(\mathbf{c}_{i'}\) if:
9.2. Exercise 1:#
Implement a function that generates points from Gaussian distributions. The function should take the following parameters:
def generar_datos_gaussianos(n_muestras=100, n_clusters=3, centros=None):
"""
Genera datos sintéticos siguiendo distribuciones gaussianas de forma simplificada.
Parámetros:
-----------
n_puntos_por_cluster : int
Cantidad de puntos para cada cluster
n_clusters : int
Número de clusters a generar
centros : array o None
Coordenadas de los centros (x,y).
Retorna:
--------
X : array
Coordenadas (x,y) de todos los puntos
y : array
Etiqueta del cluster al que pertenece cada punto
"""
return X, y
After generating the data, plot the points using different colors for each cluster. And the result should be like this:
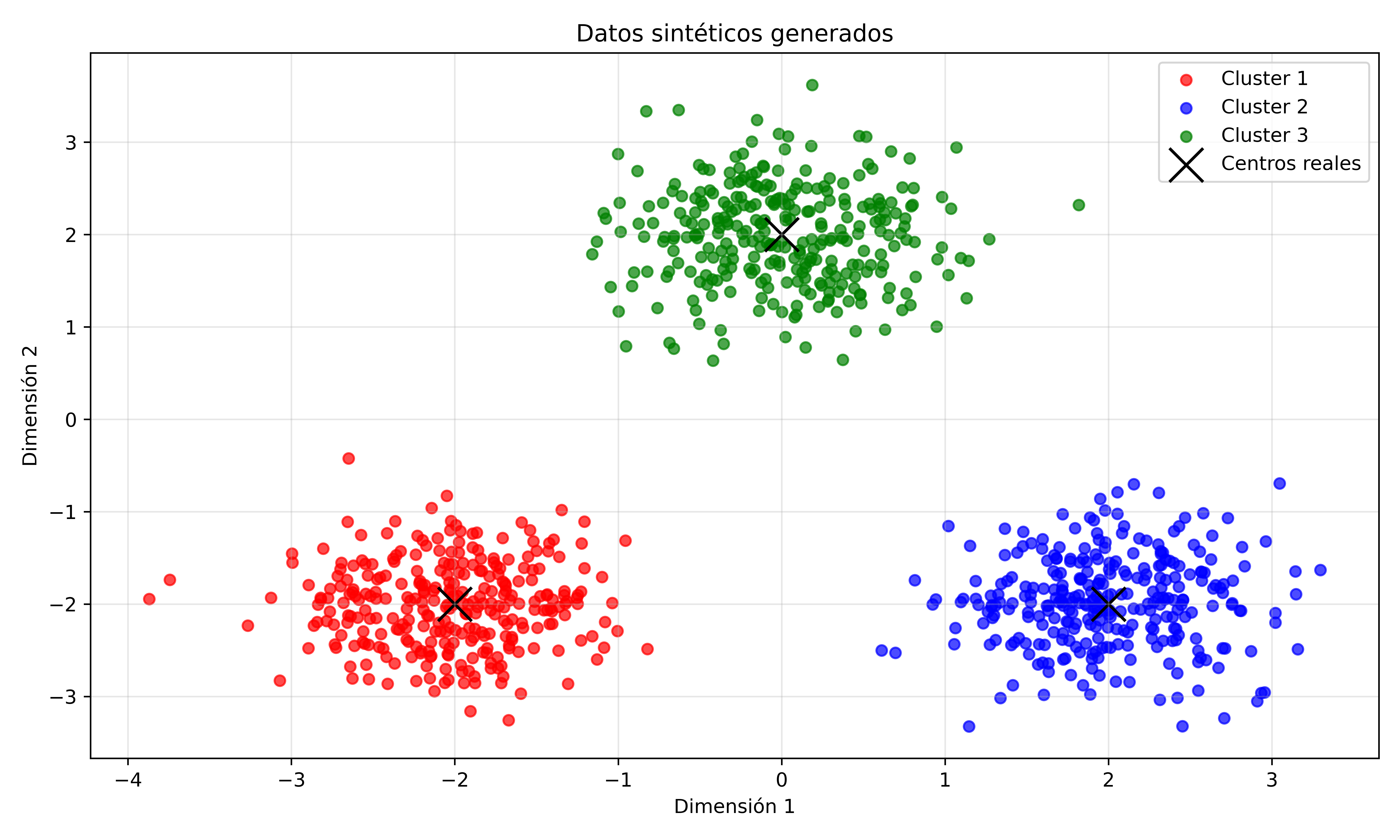
Fig. 9.1 Visualización de datos generados con distribuciones gaussianas#
9.3. Exercise 2: Implementing Deterministic Annealing#
Implement the Deterministic Annealing algorithm for clustering. You can use the following template:
import numpy as np
class DeterministicAnnealing:
def __init__(self, n_clusters=3, T_min=0.01, T_max=5.0, epsilon=1e-6, max_iter=100):
"""
Implementación del algoritmo Deterministic Annealing para clustering.
Parámetros:
-----------
n_clusters : int
Número de clusters a encontrar
T_min : float
Temperatura mínima para detener el enfriamiento
T_max : float
Temperatura inicial
epsilon : float
Umbral de convergencia para la matriz de pertenencia
max_iter : int
Número máximo de iteraciones
"""
self.n_clusters = n_clusters
self.T_min = T_min
self.T_max = T_max
self.epsilon = epsilon
self.max_iter = max_iter
self.centroids = None
self.M = None # Matriz de pertenencia
def _calcular_distancia(self, X, centroides):
"""
Calcula la matriz de distancias euclidianas al cuadrado entre puntos y centroides.
Utilizamos la identidad: ||a-b||² = ||a||² + ||b||² - 2<a,b>
donde <a,b> es el producto escalar.
"""
# Evitar errores numéricos
return np.maximum(distancias, 0)
def _actualizar_pertenencia(self, X, centroides, T):
"""
Actualiza la matriz de pertenencia (probabilidades).
Esta es la parte clave del algoritmo deterministic annealing:
la probabilidad de pertenencia depende de la distancia y la temperatura.
"""
return M
def _actualizar_centroides(self, X, M):
"""
Actualiza las posiciones de los centroides basado en la matriz de pertenencia.
Cada centroide se actualiza como un promedio ponderado de todos los puntos,
donde los pesos son las probabilidades de pertenencia.
"""
def fit(self, X):
"""
Ajusta el modelo de Deterministic Annealing a los datos.
Proceso:
1. Inicializar centroides y temperatura
2. Bucle principal de annealing:
- Calcular matriz de pertenencia
- Actualizar centroides
- Reducir temperatura
- Verificar convergencia
"""
n_muestras, n_dimensiones = X.shape
# Iniciamos con el bariocentro de los datos
self.centroids = np.mean(X, axis=0, keepdims=True)
self.centroids = np.repeat(self.centroids, self.n_clusters, axis=0)
# Inicializar parámetros para el bucle
t = 0 # Contador de iteraciones
T = self.T_max # Temperatura inicial
convergencia = False
M_anterior = np.ones((n_muestras, self.n_clusters)) / self.n_clusters
# Bucle principal
while not convergencia:
# Actualizar matriz de pertenencia con temperatura actual
# Actualizar centroides
# Actualizar contador y temperatura
# Verificar convergencia
# Guardar matriz actual para próxima iteración
return self
def predict(self, X):
"""
Predice el cluster para cada punto en X.
Retorna el índice del cluster con mayor probabilidad para cada punto.
"""
def fit_predict(self, X):
"""
Ajusta el modelo y predice los clusters en un solo paso.
"""
9.4. Exercise 3: Testing the Algorithm#
Generate synthetic data using the function implemented in Exercise 1 and apply the Deterministic Annealing algorithm to cluster the data. You should try at least the following configurations, also, it is encouraged to create more experiments to test the algorithm, with more or less dispersion, more or less clusters, etc.
Generate data with 3 clusters and apply the algorithm with 6 clusters.
Generate data with 3 clusters and apply the algorithm with 2 clusters.
Generate data with 5 clusters and apply the algorithm with 5 clusters.
Generate data with 5 clusters and apply the algorithm with 2 clusters.
Generate data with 5 clusters and apply the algorithm with 10 clusters.
9.5. Exercise 4: Entropy#
Implement a function that calculates the entropy of the clustering. The function should take the following parameters:
def calcular_entropia_por_punto(self):
"""
Calcula la entropía condicional H(C|X), que es la entropía por punto.
Esta entropía mide la incertidumbre en la asignación de clusters para cada punto.
"""
def calcular_entropia_por_cluster(self):
"""
Calcula la entropía condicional H(X|C), que es la entropía por cluster.
Esta entropía mide qué tan uniforme es la distribución de puntos dentro de cada cluster.
También devuelve un array con las entropías individuales de cada cluster.
"""
Now re-discuss the previous experiments using the entropy as a metric to evaluate the quality of the clustering. You are encouraged to create more experiments to test the algorithm. Also, you can help yourself with plots to visualize the results or how the entropy changes with the iterations of the while loop. Bonus: If you visualize the entropy per point, by changing the size of the points, making the size of the points proportional to the entropy (bigger size = higher entropy) and discuss what is happening, you will be rewarded with bonus points.
9.6. Exercise 5: Automated Clustering#
After implementing the Deterministic Annealing algorithm and the entropy function, you can now create a function that automatically clusters the data. So, now the question is: how to choose the number of clusters? The criteria will be given by the elbow method, which is a heuristic used in cluster analysis to determine the optimal number of clusters in a dataset. The elbow method involves plotting the explained variance as a function of the number of clusters and looking for the “elbow” point where the rate of decrease sharply changes.
The elbow point indicates the optimal number of clusters, as adding more clusters beyond this point does not significantly improve the explained variance.
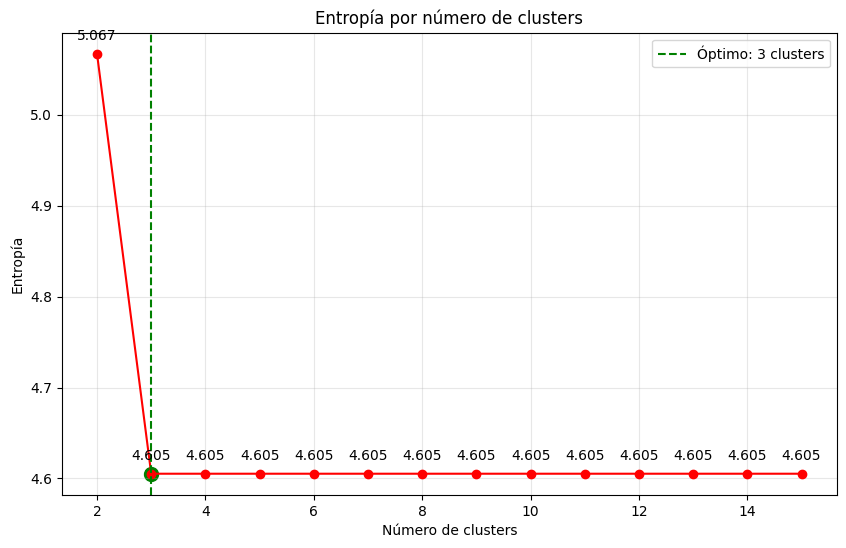
Fig. 9.2 Visualization of the entropy as a function of the number of clusters#
Here we are asking you is to explain the elbow method and how to implement it. And then, use it to find the optimal number of clusters for the data generated in Exercise 1. Obiously, you will have to discuss the results and the elbow point.
9.7. Exercise 6: Image Segmentation#
In this exercise, we will apply the Deterministic Annealing algorithm to segment an image. The goal is to cluster the pixels of the image into different segments based on their color values. We will provide you the part that handles the image loading and visualization, and you will have to implement the segmentation using the algorithm you implemented in Exercise 2.
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
from scipy import ndimage
# Fijamos las semillas para reproducibilidad
np.random.seed(42)
# 1. Cargar una imagen
image = np.array(Image.open('imagen.jpg'))
# 2. Preprocesamiento (opcional)
# Aplicar un filtro de mediana para reducir el ruido, cuando decimos ruido nos referimos a que haya demasiada variación en los colores de los píxeles
# en la imagen. Esto puede ayudar a mejorar la segmentación.
image_filtered = ndimage.median_filter(image, size=10) # ndimage.median_filter es una función de SciPy que aplica un filtro de mediana a la imagen. Le pasamos la imagen y el tamaño del filtro (10x10 píxeles en este caso). Esto significa que cada píxel en la imagen filtrada es reemplazado por la mediana de los píxeles en una vecindad de 10x10 píxeles alrededor de él.
# 2.1 Mostramos la imagen original y la imagen filtrada
plt.figure(figsize=(10, 5))
plt.subplot(121), plt.imshow(image), plt.title('Original')
plt.subplot(122), plt.imshow(image_filtered), plt.title('Filtrada')
plt.show()
# 3. Reformatear la imagen para clustering
print("Shape de la imagen original:", image.shape)
pixel_values = image_filtered.reshape((-1, 3))
print("Shape de la imagen reformateada:", pixel_values.shape)
pixel_values = np.float32(pixel_values)
# 4. Aplicar Deterministic Annealing
# ...
# 5. Visualizar los resultados
plt.figure(figsize=(15, 5))
plt.subplot(131), plt.imshow(image), plt.title('Original')
plt.subplot(132), plt.imshow(segmented_image), plt.title('Segmentada con DA')
plt.show()
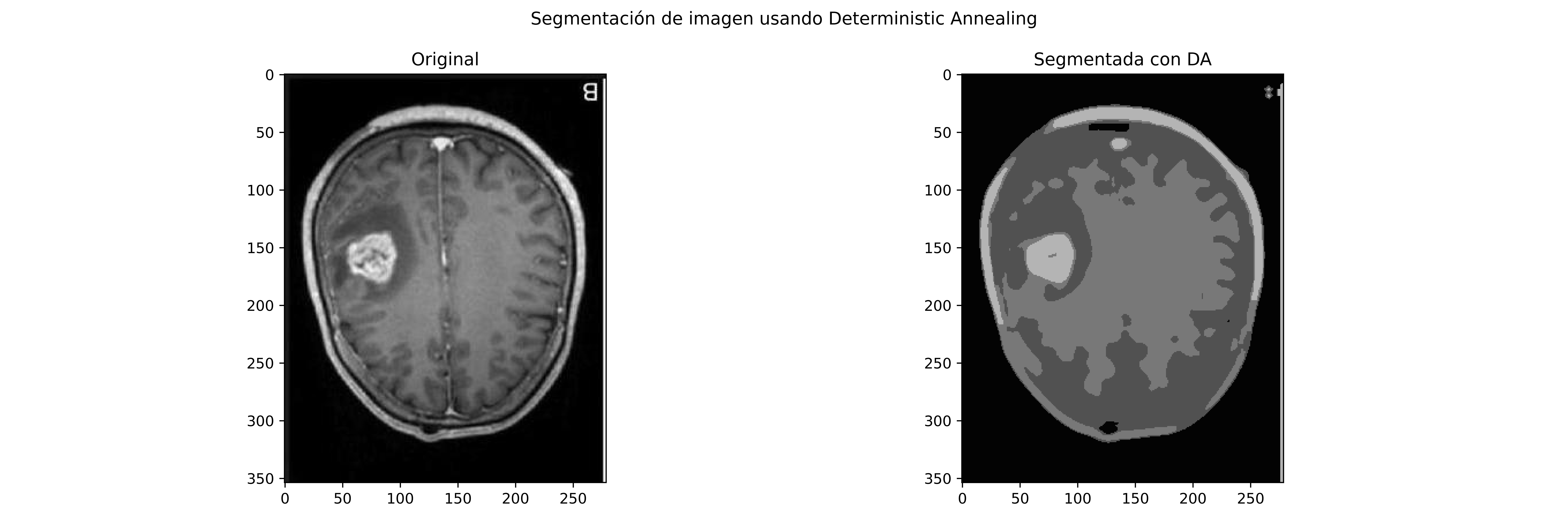
Fig. 9.3 Segmentación de la imagen utilizando el algoritmo de Deterministic Annealing#
